Simulating Lipid Vesicles
- Details
- Last Updated: Wednesday, 10 February 2021 13:14
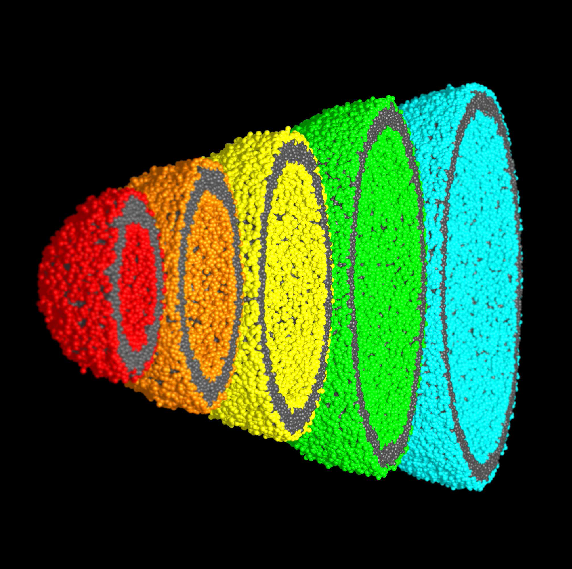
Liposomes, or vesicles, are widely used in in vitro studies as mimics of either complete cells or cell organelles such as endosomes or transport vesicles involved in endo- or exocytosis and protein trafficking. In the realm of synthetic biology they furthermore play an important role as drug carriers, sensors and many more potential applications only limited by imagination.
The coarse grained Martini model allows us to study small vesicles at near atomic detail. In an ongoing series of projects we and others have looked at the self-assembly of these vesicles [3,4], the effect of the curved geometry on lipid packing and transmonolayer lipid distribution [2,4], the fusion of vesicles [5], the freezing process [1], and structural properties of Lo/Ld phase separated vesicles [7,8,9].
The effect of a hypo-osmotic shock on a vesicle has also been simulated, both for pure lipid vesicles and for vesicles equipped with a mechano-sensitive channel [6]. A vesicle model for an entire endosome has been used in [11].
For giant vesicles, you can try the Dry Martini version [10], and for generating the starting structures of vesicles or using multiresolution models, you may try the methods described in [12-14].
- [1] H.J. Risselada, S.J. Marrink. The freezing process of small lipid vesicles at molecular resolution. Soft Matter, 5:4531-4541, 2009.
- [2] H.J. Risselada, S.J. Marrink. Curvature effects on lipid packing and dynamics in liposomes revealed by coarse grained molecular dynamics simulations. Phys. Chem. Chem. Phys., 11:2056-2067, 2009.
- [3] H.J. Risselada, A.E. Mark, S.J. Marrink. The application of mean field boundary potentials in simulations of lipid vesicles. JPC-B, 112:7438-7447, 2008.
- [4] S.J. Marrink, A.E. Mark. Molecular dynamics simulation of the formation, structure, and dynamics of small phospholipid vesicles. JACS, 125:15233-15242, 2003.
- [5] S.J. Marrink, A.E. Mark. The mechanism of vesicle fusion as revealed by molecular dynamics simulations. JACS, 125:11144-11145, 2003.
- [6] M. Louhivuori, H.J. Risselada, E. van der Giessen, S.J. Marrink. Release of content through mechano-sensitive gates in pressurized liposomes. PNAS, 107:19856-19860, 2010. open access
- [7] H.J. Risselada, S.J. Marrink.The molecular face of lipid rafts in model membranes. PNAS, 105:17367-17372, 2008.open access
- [8] H.J. Risselada, S.J. Marrink, M. Muller. Curvature-dependent elastic properties of liquid-ordered domains result in inverted domain sorting on uniaxially compressed vesicles. Phys. Rev. Lett., 106:148102, 2011. abstract
- [9] C.A. Lopez, A.H. de Vries, S.J. Marrink. Computational microscopy of cyclodextrin mediated cholesterol extraction from lipid model membranes. Sci. Rep., 3:2071, 2013. open access
- [10] C.. Arnarez, J.J. Uusitalo, M.F. Masman, H.I. Ingólfsson, D.H. de Jong, M.N. Melo, X. Periole, A.H. De Vries, S.J. Marrink. Dry Martini, a coarse-grained force field for lipid membrane simulations with implicit solvent. JCTC, 11:260–275, 2015. abstract
- [11] B.M.H. Bruininks, P.C.T. Souza, H. Ingolfsson, S.J. Marrink. A molecular view on the escape of lipoplexed DNA from the endosome. eLife, 9:e52012, 2020. doi.org/10.7554/eLife.52012
- [12] Y. Qi, H.I. Ingólfsson, X. Cheng, J. Lee, S.J. Marrink, W. Im. CHARMM-GUI Martini Maker for coarse-grained simulations with the Martini force field. JCTC, 11:4486–4494, 2015. abstract
- [13] Y. Liu, A.H. de Vries, J. Barnoud, W. Pezeshkian, J. Melcr, S.J. Marrink. Dual resolution membrane simulations using virtual sites. JPCB 124:3944, 2020. doi.org/10.1021/acs.jpcb.0c01842
- [14] W. Pezeshkian, M. Konig, T.A. Wassenaar, S.J. Marrink. Backmapping triangulated surfaces to coarse-grained membrane models. Nature Commun. 11:2296, 2020. doi.org/10.1038/s41467-020-16094-y


























































































































































