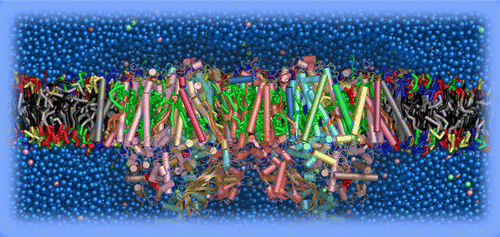
Polarizable water model
- Details
- Last Updated: Monday, 18 September 2017 10:39
A refined polarizable water model is now available, developed by the group of Jens Smiatek. The new model improves the behavior of Martini with long range electrostatic interactions. Available from the download section. See: J. Michalowsky, L.V. Schäfer, C. Holm, J. Smiatek. http://dx.doi.org/10.1063/1.4974833 for details.